Genotype
This article has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
(Learn how and when to remove this template message)
|
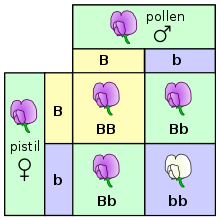
The genotype is the part of the genetic makeup of a cell, and therefore of any individual, which determines one of its characteristics (phenotype).[1] The term was coined by the Danish botanist, plant physiologist and geneticist Wilhelm Johannsen in 1903.[2]
Genotype is one of three factors that determine phenotype, along with inherited epigenetic factors and non-inherited environmental factors. Not all organisms with the same genotype look or act the same way because appearance and behavior are modified by environmental and growing conditions. Likewise, not all organisms that look alike necessarily have the same genotype.
One's genotype differs subtly from one's genomic sequence, because it refers to how an individual differs or is specialized within a group of individuals or a species. So, typically, one refers to an individual's genotype with regard to a particular gene of interest and the combination of alleles the individual carries (see homozygous, heterozygous).[3] Genotypes are often denoted with letters, for example Bb, where B stands for one allele and b for another.
Somatic mutations which are acquired rather than inherited, such as those in cancers, are not part of the individual's genotype. Hence, scientists and physicians sometimes talk about the genotype of a particular cancer, that is, of the disease as distinct from the diseased.
An example of a characteristic determined by a genotype is the petal color in a pea plant. The collection of all genetic possibilities for a single trait are called alleles; two alleles for petal color are purple and white.[4]
Contents
Phenotype[edit]
Any given gene will usually cause an observable change in an organism, known as the phenotype. The terms genotype and phenotype are distinct for at least two reasons:
- To distinguish the source of an observer's knowledge (one can know about genotype by observing DNA; one can know about phenotype by observing outward appearance of an organism).
- Genotype and phenotype are not always directly correlated. Some genes only express a given phenotype in certain environmental conditions. Conversely, some phenotypes could be the result of multiple genotypes. The genotype is commonly mixed up with the phenotype which describes the end result of both the genetic and the environmental factors giving the observed expression (e.g. blue eyes, hair color, or various hereditary diseases).
A simple example to illustrate genotype as distinct from phenotype is the flower colour in pea plants (see Gregor Mendel). There are three available genotypes, PP (homozygous dominant), Pp (heterozygous), and pp (homozygous recessive). All three have different genotypes but the first two have the same phenotype (purple) as distinct from the third (white).
A more technical example to illustrate genotype is the single-nucleotide polymorphism or SNP. A SNP occurs when corresponding sequences of DNA from different individuals differ at one DNA base, for example where the sequence AAGCCTA changes to AAGCTTA. This contains two alleles : C and T. SNPs typically have three genotypes, denoted generically AA Aa and aa. In the example above, the three genotypes would be CC, CT and TT. Other types of genetic marker, such as microsatellites, can have more than two alleles, and thus many different genotypes.
Penetrance is the proportion of individuals showing a specified genotype in their phenotype under a given set of environmental conditions.[5]
Mendelian inheritance[edit]
The distinction between genotype and phenotype is commonly experienced when studying family patterns for certain hereditary diseases or conditions, for example, hemophilia. Humans and most animals are diploid; thus there are two alleles for any given gene. These alleles can be the same (homozygous) or different (heterozygous), depending on the individual (see zygote). With a dominant allele, the offspring is guaranteed to inherit the trait in question irrespective of the second allele.
In the case of an albino with a recessive allele (aa), the phenotype depends upon the other allele (Aa, aA, aa or AA). An affected person mating with a heterozygous individual (Aa or aA, also carrier) there is a 50-50 chance the offspring will be albino's phenotype. If a heterozygote mates with another heterozygote, there is 75% chance passing the gene on and only a 25% chance that the gene will be displayed. A homozygous dominant (AA) individual has a normal phenotype and no risk of abnormal offspring. A homozygous recessive individual has an abnormal phenotype and is guaranteed to pass the abnormal gene onto offspring.
In the case of hemophilia, it is sex-linked thus only carried on the X chromosome. Only females can be a carrier in which the abnormality is not displayed. This woman has a normal phenotype, but runs a 50-50 chance, with an unaffected partner, of passing her abnormal gene on to her offspring. If she mated with a man with haemophilia (another carrier) there would be a 75% chance of passing on the gene.
Determination[edit]
Genotyping is the process of elucidating the genotype of an individual with a biological assay. Also known as a genotypic assay, techniques include PCR, DNA fragment analysis, allele specific oligonucleotide (ASO) probes, DNA sequencing, and nucleic acid hybridization to DNA microarrays or beads. Several common genotyping techniques include restriction fragment length polymorphism (RFLP), terminal restriction fragment length polymorphism (t-RFLP),[6] amplified fragment length polymorphism (AFLP),[7] and multiplex ligation-dependent probe amplification (MLPA).[8]
DNA fragment analysis can also be used to determine such disease causing genetics aberrations as microsatellite instability (MSI),[9] trisomy[10] or aneuploidy, and loss of heterozygosity (LOH).[11] MSI and LOH in particular have been associated with cancer cell genotypes for colon, breast and cervical cancer.
The most common chromosomal aneuploidy is a trisomy of chromosome 21 which manifests itself as Down syndrome. Current technological limitations typically allow only a fraction of an individual's genotype to be determined efficiently.
See also[edit]
- Endophenotype
- Genotype–phenotype distinction
- Nucleic acid sequence
- Phenotype
- Potentiality and actuality
- Quaternary numeral system
- Sequence (biology)
References[edit]
- ^ "Genotype definition - Medical Dictionary definitions". medterms.com.
- ^ Johannsen, W. (1903). 'Om arvelighed i samfund og i rene linier', Oversigt over det Kongelige Danske Videnskabernes Selskabs Forhandlinger, vol. 3, 247-70. German ed. Erblichkeit in Populationen und in reinen Linien. Jena: Gustav Fischer, 1903; Scanned full text. Also see his monograph Arvelighedslærens elementer ("The Elements of Heredity". Copenhagen, 1905), which was rewritten, enlarged and translated into German as Elemente der exakten Erblichkeitslehre (Jena: Gustav Fischer, 1905; Scanned full text.
- ^ Griffiths AJF, Gelbart WM, Miller JH, et al. Modern Genetic Analysis. New York: W. H. Freeman; 1999. Genetics begins with Variation. Available from: https://www.ncbi.nlm.nih.gov/books/NBK21344/
- ^ Alberts, Bruce; Bray, Dennis; Hopkin, Karen; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter (2014). Essential Cell Biology (4 ed.). New York, NY: Garland Science. p. 659. ISBN 978-0-8153-4454-4.
- ^ Allaby, Michael, ed. (2009). A dictionary of zoology (3rd ed.). Oxford: Oxford University Press. ISBN 9780199233410. OCLC 260204631.
- ^ Hulce, David; Liu, ChangSheng (Jonathan) (July 2006). "SoftGenetics Application Note - GeneMarker® Software for Terminal-Restriction Fragment Length Polymorphism (T-RFLP) Data Analysis" (PDF). SoftGenetics. Archived from the original (PDF) on 2007-06-13.CS1 maint: Multiple names: authors list (link)
- ^ "Keygene.com Homepage" Archived 2011-06-28 at the Wayback Machine
- ^ "SoftGenetics Application Note - Software for Multiplex Ligation-dependent Probe Amplification (MLPA™)" (PDF). SoftGenetics. April 2006. Archived from the original (PDF) on 2011-07-16. Retrieved 2011-03-13.
- ^ He, Haiguo; Ning, Wan; Liu, Jonathan (March 2007). "SoftGenetics Application Note - Microsatellite Instability Analysis with GeneMarker® Tamela Serensits" (PDF). SoftGenetics. Archived from the original (PDF) on 2007-09-23.
- ^ "SoftGenetics Application Note - GeneMarker® Software for Trisomy Analysis" (PDF). SoftGenetics. November 2006. Archived from the original (PDF) on 2007-07-28.
- ^ Serensits, Pamela; He, Haiguo, Ning, Wan; Liu, Jonathan (March 2007). "SoftGenetics Application Note - Loss of Heterozygosity Detection with GeneMarker" (PDF). SoftGenetics. Archived from the original (PDF) on 2007-07-28.CS1 maint: Multiple names: authors list (link)
External links[edit]
| Look up genotype, phenotype, inheritance, or genome in Wiktionary, the free dictionary. |
| Wikimedia Commons has media related to Genotypes. |